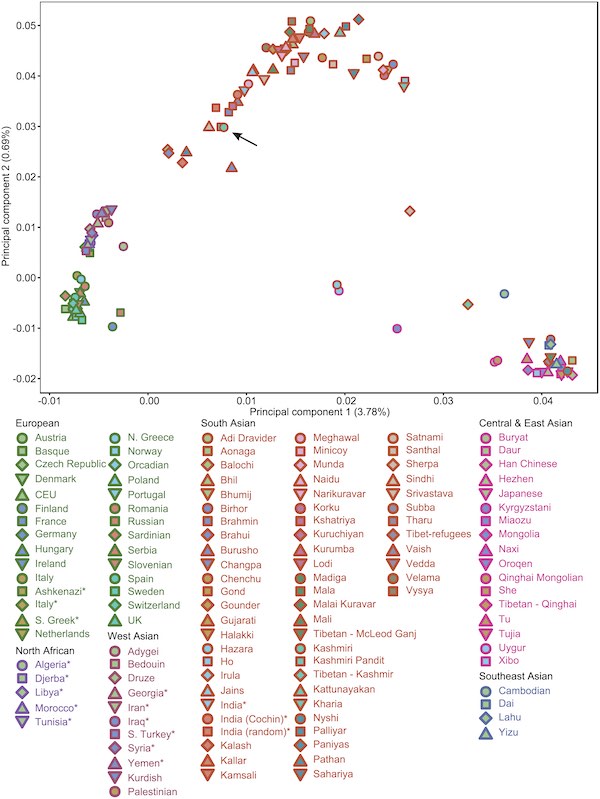
I think I’ve posted this before, but it was a while ago before we had so many readers. In this paper they took 15 random Kashmiris from the Valley, and compared them to various populations. The plot below, as well as admixture analysis in the paper, shows no daylight between the Pandit samples and generic Muslim Kashmiris.
This is not to say Pandits are not an endogamous community and were not before the Islamicization of Kashmir. But, it is to say that in their overall genome their origins are exactly the same as other Kashmiris. This is in contrast to many parts of India in regards to Brahmins, though the “stylized fact” seems to be the further north and west you go, the smaller the genome-wide difference between Brahmins and non-Brahmins will be. This seems to comport with the idea that Brahmins are intrusive to the south and east in a way they are not to the north and west.
Finally, the data from ancient DNA is strongly suggestive of “AASI-reflux” across north and west South Asia after 3000 BC. See my post The Aryan Integration Theory (AIT).

This might be a good time to ask a question that’s been bothering me about the statistical interpretation of human genetics. In order to have an uncertainty of 1 sigma in a result, one would need to randomly sample the the square root of N, out of a total population of N. In some fields, a conclusive statement requires much more accuracy. For example, in particle physics we require a 5 sigma accuracy to claim a discovery. Is there a consistent standard which is followed in population genetics? I’m asking this because a sample of 15 Kashmiri Pandits doesn’t seem sufficient to make the kind of strong statement that you just made. Also, if I may add, the same things bugs me when I look at the analysis of Riech’s lab. Ancient India had a population of roughly 4 million at any given time. In order to make a definitive statement about the genome of Ancient Indians over a period of 500 years from 2000 to 1500 BC, which is the relevant time frame for AIT, AMT, and the rest, we are looking at roughly 20 generations. That makes N of the order 100 million. Naively, I would have thought that in order to make any claims regarding AIT, AMT, OIT, one would then require a sample of at least 10,000 people. Moreover, the historical samples we might come across wouldn’t be random so we would need an even larger sample and then statistically correct for survival bias. But the sample size of most of these studies is an order of magnitude less than that. What’s going on here?
Is there a consistent standard which is followed in population genetics?
no, because the effective population size and nature of population structure varies a lot btwn species/groups. you kind of follow heuristics based on your lineage being sampled.
the census sizes for humans are not informative. what you want is the longer term effective pop size (harmonic mean of census) and so the bottleneck is what really matters. that’s how the ‘long term effective pop size’ of non-african humans is 1,000 even though our census size today is like 7 billion. the mutation rate isn’t high enough to replenish all the variation that was removed through this bottleneck 50,000 years ago.
a second issue is that a single individual has within their whole genome (say 50 million polymorphic markers out of 3 billion base pairs) the phylogeny of all their ancestors. when you sample a single individual and get genome-wide information you aren’t really just an N = 1. that’s how you can infer the whole population history of non-africans from a single whole genome (google PSMC).
the more structure and within-group homogeneity you have the less likely you are going to get structure-informative variance out of more samples. south asians are very structured, so when i get a sample from a clear and distinct jati that tells me most of what i know. more than 5 is really not useful for the sort of questions mooted on this weblog (it is useful for stuff like low frequency private variants). there is also really really subtle structure ppl have found in iyers and Ashkenazi jews, but this is really on-the-margin stuff. really when you are sampling from an endogamous homogeneous group you are getting more information on the same genealogy over and over.
tldr; i would be surprised if there were some minority groups in kashmir that plot in a different place on the PCA. but i think probably not since the random Muslim and pandits are on the same spot.
make sense?
Thanks for reply. The second point about a single individual being not just N=1 due to them carrying millions of genetic markers makes sense. I would have to read up a bit more on the effective pop size to truly understand the first point.
>>the more structure and within-group homogeneity you have the less likely you are going to get structure-informative variance out of more samples. south asians are very structured, so when i get a sample from a clear and distinct jati that tells me most of what i know.
I understand this point. Although my doubt is that in order to establish the fact that a particular population is structured, we would have to use a much larger sample and once it is established that it is structured, we can then use a small sample to get the important information that you are referring to. Similarly, I would have thought that in order to estimate what the effective pop size is, we would have to consider a statistically meaningful sample of the census(of the order of square root of N). And once we have the effective pop size, then we needn’t worry about the large sample. Likewise, I would think that in order to establish the fact that a particular jaati is endogamous a much larger sample is needed and once that’s established, we can use, like you said, N=5 to compare the genetic differences between two jatis. Similarly, my naive guess would have been that the sample required to predict AIT or OIT would be of the order the square root of N since we are making a statement without any prior knowledge of the migratory patterns of a population. Once it is established, then we can just use a few Indians and a few central Asians or Europeans to compare the similarities or the differences. Or am I not understanding the concept of an effective pop size correctly?
Green Veggies, that’s because each “sample” is really 10,000 samples. Every person is descended from thousands of people and contains samples from all of them. The amount of information in one genome is gigantic. The only real issue is if your only sample is an outlier like the Uralic people you find among the Sintashta graves or the Altaics among the Afanaseivo graves.
Thanks, that makes sense. Presmably that is what Razib was trying to explain as well.
Just look at the surnames of Some Muslims Kashmiris for instance like Riyaz Naikoo, Afzal Guru , Bhatt etc.
But isn’t that true that some people migrated from Bukhara or from Iran during medieval times.
I think so. I score some Iranian Mazandarani on my global25 but I’m also a Kashmiri Pandit. I don’t know how prevalent inter marriage was but I do know I have an indirect ancestor who married an Iranian woman, plus a few other recent relatives who intermarried with Kashmiri Muslims.
But isn’t that true that some people migrated from Bukhara or from Iran during medieval times.
they seem pretty generic north indians but i will try and look deeper
Could you please let me know which y dna is common among Kashmiri Muslims n pandits thanks
But Kashmiri Hindus have much more common with Muslims in terms 9f ethnicity,food,dressing( phiran i know about) etc. But eventually civilizationally there are connected to India.
Well, the food and dress is more of a climatic necessity. The pheran was a means to keep warm (layered and made of wool in winter and also loose enough one could fit a kangri inside). Food, yes we are meat eaters traditionally but this isn’t unique only to Kashmiri pandits as I believe bengali brahmins and certain other groups of Brahmins do eat meat. Kashmiri Pandit cuisine traditionally omits onions and garlic, too. There’s also been an adoption of things like vegetarianism and celebration of Karva Chauth and Diwali amongst Kashmiri pandits as a way to identify with our faith more since there is a perception we are more similar to Kashmiri Muslims rather than other Hindus.
@Razib
I had a question to ask regarding the steppe migrations. How possible it is to have multiple migration events separated by long time interval. I have read from your writings that brahmins cannot be modeled to be derived from single group which points to multiple IA groups entering South asia. But the migration event as whole is still seen around 1500 BC from northwest.
I read the discussion around using MLBA or EMBA as source population for south asia or R1a L657 presence instead of R1a Z93 in South asia. Can the first arrival of R1a be pushed back in 3rd millennium using path through tibet bypassing BMAC/ IVC (see image). I could also add point like mount kailash, eastern uttarakhand to this route since many holy sites of hinduism exist in that area.
https://imgur.com/TQAEH50
Old parts of Rgveda can be attributed to this group mixed with AASI since the setting is non urban/ forest but still in South aisa. Later the second wave of steppe through northwest as mentioned in Narasimhan paper results in mixing of IVC, AASI , earlier and later steppe inputs as function of geography.
In this scenario the steppe ancestry for Kalash and bhumihar_bihar is equally high but from different sources as tabulated in Aryan integration theory post of yours. The kalash therefore no longer represent ideal ANI proxy for whole of South Asia. This could also explain lack of dravidian toponyms in gangetic plain as this area was occupied by IA much before the post IVC mixing.
Question based on some conversations on this blog and limited reading on genetics and therefore could be completely wrong for having overlooked basic facts. Thanks in advance.
What does this, if any speculation is possible, imply about the history of caste in the valley?
The Central Asian component in Kashmir, apart from the generic one in North Indians, comes from the few migrants from the area in recent centuries. This category stands apart from other Kashmiri Muslims by their surnames,-thus Bokhari, Hamdani, Shah, Naqshbandi, Mufti, Geelani etc, as compared to direct Pandit lineages, or of other classes such as Dars, Tantrey, Parray, Khandey. The imported (for want of a better term) lineage is referred to as Malla (distinct from Mallah, boatman) by other Kashmiri Muslims. The Mulla intermarry among themselves generally, and are accused of clannishness and nepotism. They are mostly confined to Srinagar city, and are the descendants of attendants to the various religious Preceptors who migrated to the valley from Iran and Central Asia, in particular.Shah Hamdan, known popularly as Mir Kabir. Shah Hamdani is said to have been accompanied by 700 Syeds, who presumably stayed back when Shah Hamdan left. There are also a few direct descendants of Afghans, but these have generally merged with native Kashmiris through intermarriage in recent decades, as have Punjabi Muslims. There is only one large village, Guthlibag, with a substantive Afghan population where the residents still speak Pushto and marry only among themselves. Smaller enclaves are reported to exist elsewhere in South Kashmir but none came to my notice.
SAS Geelani, by the way, is neither a Syed, nor a Geelani. He was plain Ali Shah before adorning the AS with honorific prefix S and suffix G.