Earlier this month I was part of a podcast discussion about AIT and its counter (OIT?) with Razib, Mukunda, Kushal Mehra, and a Carvaka regular Kartik Mohan. It was a good discussion though I doubt if it would come out as a great podcast.
Personally, I have gone down the AIT/OIT rabbit hole enough last few years to want a long break from these discussions. However, before I take the break, I would like to summarize my current position which would also act as my notes for the podcast. For my position on this topic a year ago please find the following blogpost – From OIT to AIT
You can listen to the podcast here.
Language & Genes:
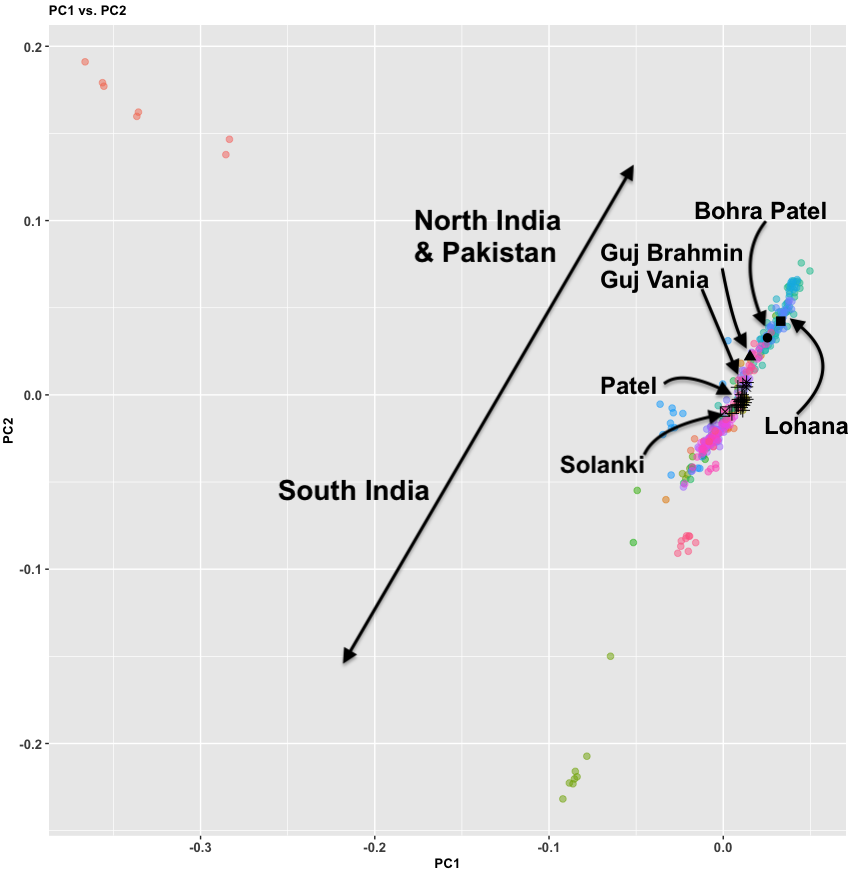
I firmly believe that ancient genetics is the strongest method for unraveling the mysteries of prehistory. For pre-modern societies, I do not believe there were any mechanisms of the spread of primary languages without mass movements of people. Language is a meme but unlike religions, it has complex mechanisms of spread that take years and requires (in most cases) familial teaching. If we take examples of memetic spread in recorded history – be it the spread of Islam in Southeast Asia or Buddhism in East Asia(via trade, etc) both happened without fundamental alterations in the primary language of the recipient regions. So in essence I do not find the model of primary language shift through mechanisms of trade or other N mechanisms posited as feasible.
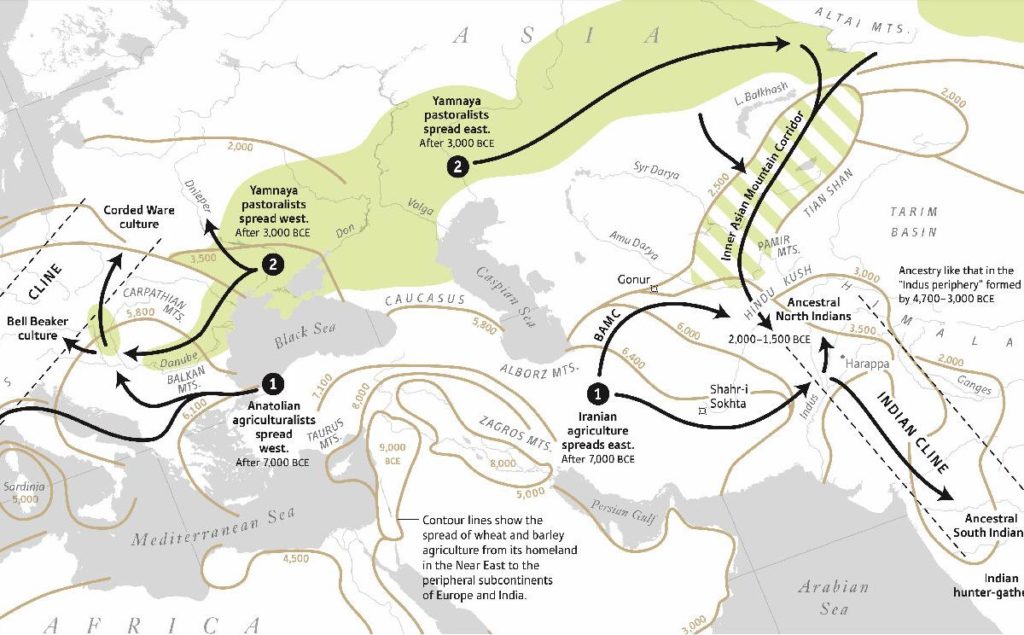
As Razib pointed out in the podcast – everywhere on the earth where Indo-European languages were spoken as primary languages in pre-modern times, the Steppe genetic signal is present in substantial amounts. I do not think any other explanation other than some form of Steppe hypothesis can explain this data. Of course, PIE homeland (including Hittite) could be in the Steppe or it could be elsewhere. I believe what we can firmly state is not where PIE originated, but where PIE developed and was spread out of.
If we want to look at a potential model of IA spread into India from the historic record we can take a look at the British isles. The languages are nicely split in the east-west direction along Germanic/Celtic lines similar to the North-South split of Aryan and Dravidian languages in India. Not only that only around 38% ancestry of Britain is Anglo Saxon with an east-west gradient.
Critics of OIT like to attack particulars of the 2007 Anthony book as new evidence is unearthed as if any dissonance in Anthony/Mallory model is a slamdunk against the Steppe hypothesis. Personally, I have no strong positions on the details of the Steppe hypothesis as argued by Anthony and Mallory. The evidence of horseback riding for the Sredny Stog or even the Yamnaya is circumstantial at best (I believe some form of horseback riding as a tool of herding by young shepherds might have been a possibility), so we cannot firmly assume that horseback riding was the reason for the massive demographic changes brought about by the Steppe men. This might have mattered before the ancient DNA revolution, but now we know the genes of the steppe pastoralists spread, maybe because of the horseback riding or maybe just due to the benefits of horse husbandry or some other reasons. We know massive demographic and (probably) linguistic changes occurred from the Steppes, the “how” question might eventually get solved (or it might not) – but that doesn’t poke holes in the larger “Steppe hypothesis” built on top of Archaeogenetics.
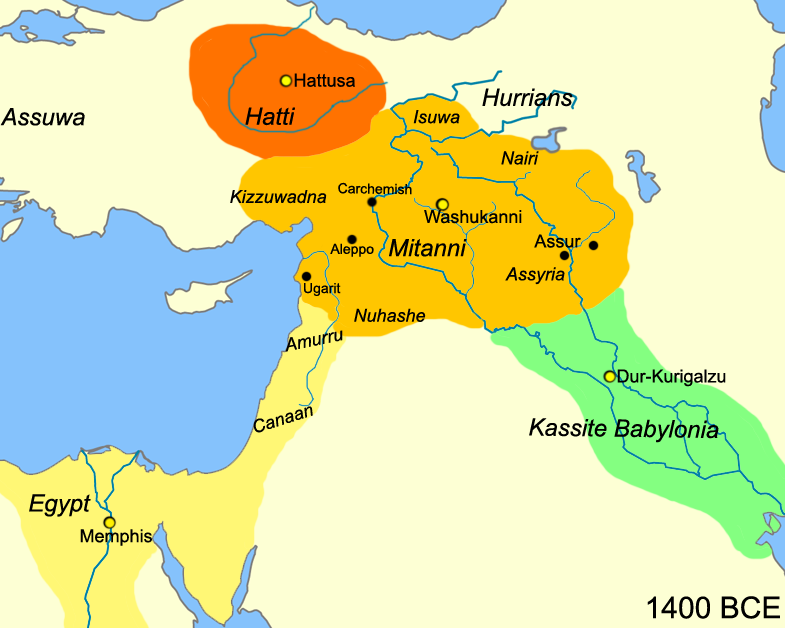
This doesn’t mean that any other mechanism of IE spread cannot work with the available data but it has to go beyond the tenuous mechanisms like trade-aided language spread. Even the Elite migration hypothesis (minus demographic changes) doesn’t seem to work as well as we assumed before the ancient DNA. The Hittite and Mitanni IE elites did not cause any substantial demographic changes nor did they cause any long-term linguistic alterations in the middle eastern region. Closer to home, Indians have had elite rules who spoke Greek, Iranian languages, Turkic languages, and lastly English. These massive elite dominations for centuries have only resulted in superstrates and languages like Urdu (I am not sure if we call it a creole). By the end of an efficient British Raj, not even 1% of Indians spoke English as a primary language. Thus anyone who tries to explain away IA spread out of India has to account for the 20-30% paternal ancestry (50% Y chromosomes) which seems to have changed in the other direction. This paternal ancestry matters a lot as we know the ancient Aryas were patriarchal, patrilineal, and patrifocal.
Also, it is often overlooked that AIT is just one node of the larger PIE – Steppe hypothesis. Even if details of AIT are contested how much does that matter to the PIE question? Also even if PIE shifts out of Pontic Steppe into Iran or Anatolia it would still be against any model for OIT and in favor of some form of AIT.
As I tried pointing out in the podcast, I see a lot of issues with Talageri and other OIT (anti-AIT scenarios) – I have not read Talageri’s books yet but have read his blog posts and interpretation of RV and listened to his podcasts on Carvaka. My points against those interpretations are
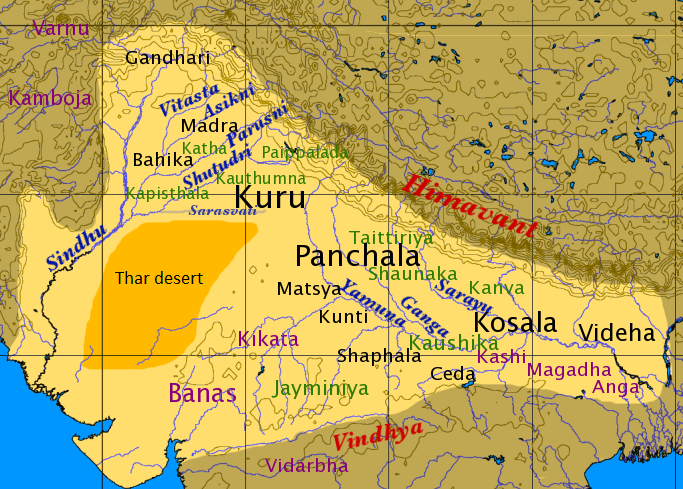
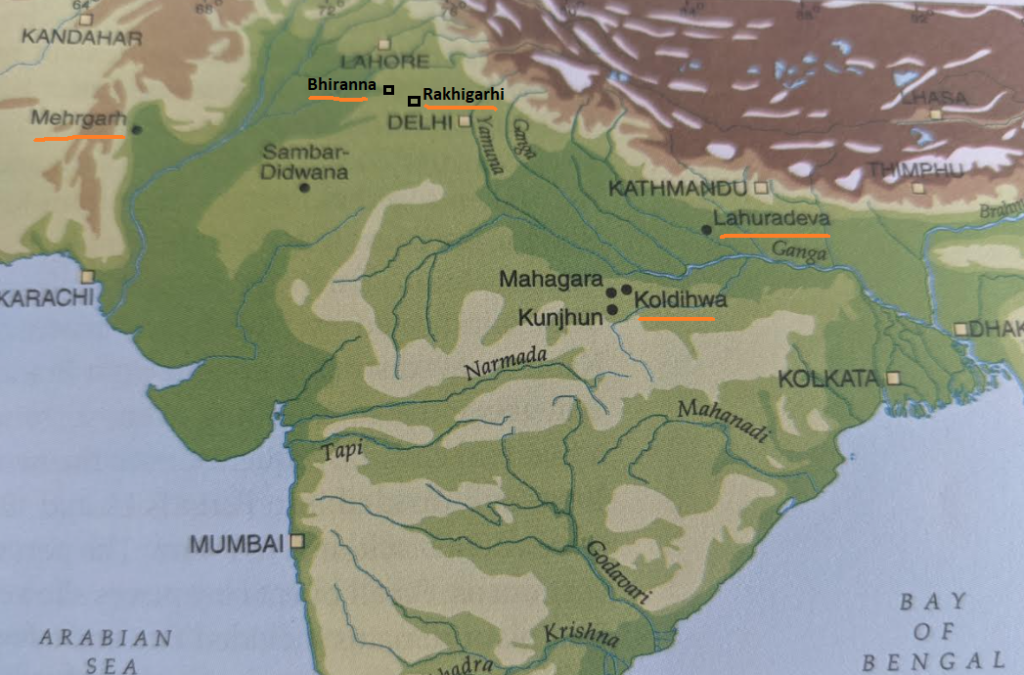
- The east to west movement of the Bharatas based on the mandalas 6-3-7 seems to hold on to some very tenuous points from the RV (eg: 2-3 references to Ganga). This reasoning might appear possible (not probable) but has zero archeological records to support it – especially for the timelines Talageri argues for (3000 BCE). Before 2000 BCE we have no archaeological data from the Gangetic plains to buttress these extraordinary claims.
- The lack of references to rice in RV is also inconsistent with the Gangetic origins of Aryas.
- The whole Asva/Ratha argument in Talageri model old RV as other equids/carts doesn’t seem to work in my preliminary reading of the RV. While the point made by Talageri “that all references to Asva/Ratha need not mean Horse/Chariot” is a correct one; the opposite isn’t automatically true => All references to Asva/Ratha need not be non Horse/non Chariot references. On the contrary, reading the RV I felt reading those references as horse/chariots make more sense (maybe it’s my priors). Interested readers can read through RV 3.43 3.45 6.29 6.44 6.45 7.18 7.19 and see for themselves if the references to Asva/Ratha appear to be for Horse/chariots or for Cart/Donkeys. (especially the Dasrajna hymns).
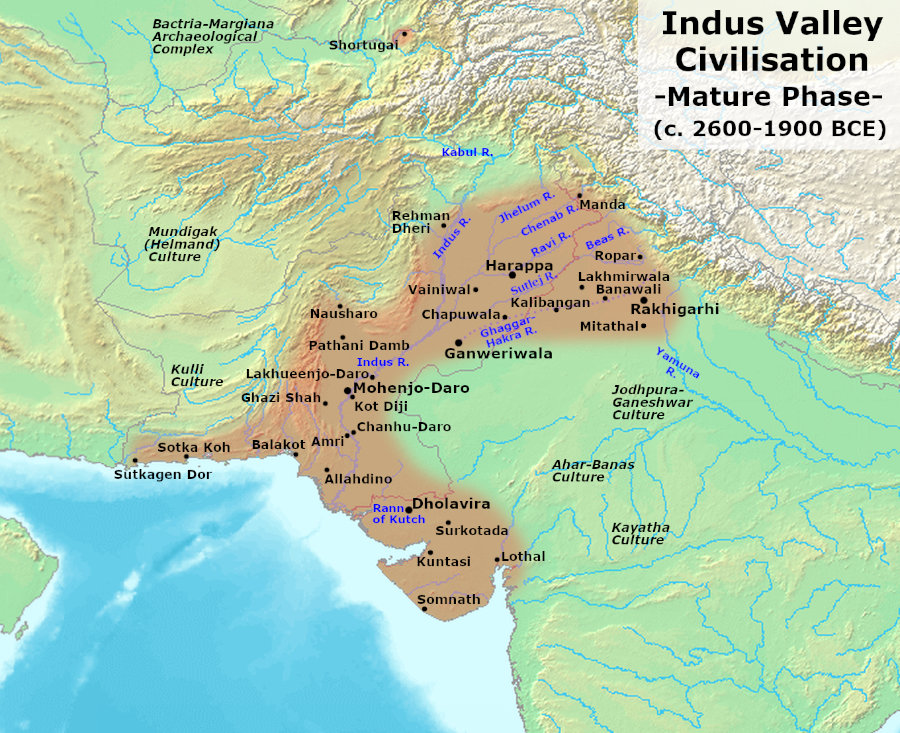
- Whatever inferences I take from RV, I find it difficult to impose them on whatever we know of the IVC. This doesn’t automatically rule out the possibility of Arya poets living on the peripheries of IVC and composing RV but makes it unlikely IMO.
However to conclusively deny these assertions one would have to do a meta-analysis of RV, if ever I get down this rabbit hole in the future I might do it myself. However in the meantime, one can look at this.
Needless to say, such interpretations will remain “circumstantial” be they in support of the AIT or OIT. After all, RV only captures a thread of the ancient Indian past while the others may be completely lost.
Personally, I would be open to alternative scenarios to explain the IA spread into India, like IA migration during the IVC (before the Sintastha) while migrations downstream of Sintastha which are attested via Genetics being responsible for the consolidation of Aryas as Kshatriya/Brahmana elites of the Vedic age. But these are extraordinary scenarios and they would require at least some robust objective pieces of evidence like
- Steppe signal from before 2000 BCE.
- Chariots or other classical IE motifs before 2000 BCE.
- Or deciphering of IVC script (or any other script from ancient India) to a Sanskrit-like language.
Outside the world of religion and mathematics, there is no absolute certainty. As a result outlier individuals from academic disciplines will continue to have non-conformist takes (like Kazanas for example). Such takes over the decades will continue to be used to create elaborate theories, be it using linguistics, genetics, and something else and they will continue getting traction in some groups (European pagans, Serb nationalists, Indian nationalists). The solutions to the PIE question are models, some more parsimonious; some tenuous, others ridiculous. There is certainly enough circumstantial data to spin wild theories putting the homeland from Iberia (initial bell beakers) to Gangetic plains (OIT), but none of these theories is the best fit for the data we have today. Maybe with newer data, some better candidates can emerge (though I doubt it). But going by the academic consensus from 3 fields -> Genetics, Linguistics, and Archaeology some model of the Steppe hypothesis is the best fit for the Indo-European question.
But for all, we know someone can still spin a theory based on some evidence that puts the PIE homeland in the sunken Atlantis.
Post Script:
I have had enough of the AIT/OIT debate and I will be avoiding this topic in the future. It has become a political and emotional topic and there is only so much that there can be no conclusion as what people assume to be at stake isn’t merely an academic question like Pre-Clovis peopling of the Americas.